Aprenda a visualizar, modelar y comprender sus proyectos medioambientales con la creación y definición de plumas de contaminación utilizando las potentes herramientas geoestadísticas de la extensión Contaminants de Leapfrog Works para realizar estimaciones transparentes y justificables de la masa y la ubicación de los contaminantes en sus proyectos.
En este seminario web se abordará lo siguiente:
- Introducción a la extensión Contaminants de Leapfrog Works.
- El flujo de trabajo de la extensión Contaminants con un breve resumen de:
- Preparación y visualización de datos
- Dominio y análisis exploratorio de datos
- Estimación
- Validación del modelo
- Informes
- Sesión de preguntas y respuestas en vivo.
Generalidades
Oradores
Aaron Hollingsworth
Especialista en soluciones para clientes en Seequent
Steve Law
Técnico sénior en Seequent
Duración
55 min
Vea más videos bajo demanda.
VideosObtenga más información acerca de la solución civil de Seequent.
Más informaciónTranscripción del video
[00:00:00.320]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />So hello everyone<encoded_tag_open />/v<encoded_tag_closed /><!– wpml:html_fragment </p> –>
<p>[00:00:01.153]<br />
and welcome to today’s webinar</p>
<p>[00:00:02.400]<br />
on the Contaminants Extension for Leapfrog Works.</p>
<p>[00:00:05.990]<br />
Today we’ll be looking at the demonstration</p>
<p>[00:00:07.530]<br />
of the Contaminant Extension</p>
<p>[00:00:09.083]<br />
with time at the end of the webinar for your questions.</p>
<p>[00:00:12.460]<br />
My name is Aaron and I’m a Customer Solution Specialist</p>
<p>[00:00:15.540]<br />
here at Seequent, based in Brisbane office</p>
<p>[00:00:17.980]<br />
and I’m joined today by Steve Law</p>
<p>[00:00:19.710]<br />
our Senior Technical Lead for Geology.</p>
<p>[00:00:21.850]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Hello, I’m Steve<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:00:22.850]<br />
and I’m a Senior Technical Lead</p>
<p>[00:00:24.520]<br />
and I’m based here in (mumbles).</p>
<p>[00:00:30.180]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Today we’ll first go through<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:00:31.280]<br />
a bit of background on Seequent</p>
<p>[00:00:32.680]<br />
and the Contaminants Extension.</p>
<p>[00:00:34.900]<br />
I’ll then hand over to Steve who will take us through</p>
<p>[00:00:37.680]<br />
the Contaminant Extension in Leapfrog Works itself.</p>
<p>[00:00:41.610]<br />
After that we’ll go through any questions that anyone has.</p>
<p>[00:00:44.240]<br />
So please if you do have any at any time,</p>
<p>[00:00:46.470]<br />
put them in here at the question or the chat window</p>
<p>[00:00:48.890]<br />
and go to webinar and we’ll get to them.</p>
<p>[00:00:53.604]<br />
There’s some background on Seequent.</p>
<p>[00:00:56.070]<br />
We are a global leader</p>
<p>[00:00:57.230]<br />
in the development of visual data science software</p>
<p>[00:00:59.470]<br />
and collaborative technologies.</p>
<p>[00:01:01.560]<br />
With our software you can turn complex data</p>
<p>[00:01:03.610]<br />
into geological understanding, providing timely insights</p>
<p>[00:01:07.220]<br />
and give decision makers confidence.</p>
<p>[00:01:10.270]<br />
At Seequent, our mission is to enable our customers</p>
<p>[00:01:12.640]<br />
to make better decisions about their earth</p>
<p>[00:01:14.590]<br />
and environmental challenges.</p>
<p>[00:01:18.430]<br />
The origins of Seequent go back to the 1990s</p>
<p>[00:01:21.570]<br />
when some smart people</p>
<p>[00:01:22.680]<br />
at the University of Canterbury in New Zealand,</p>
<p>[00:01:24.720]<br />
created the fast radio basis functions</p>
<p>[00:01:27.170]<br />
for efficiently generating 3D surfaces from point data.</p>
<p>[00:01:31.460]<br />
One of the applications that was seen for this</p>
<p>[00:01:33.300]<br />
was a better way to build geological modeling.</p>
<p>[00:01:36.080]<br />
And so in 2004, the company ARANZ Geo was founded</p>
<p>[00:01:39.490]<br />
and the Leapfrog software was first released</p>
<p>[00:01:41.680]<br />
to the mining industry.</p>
<p>[00:01:44.250]<br />
Since then the company has grown and evolved</p>
<p>[00:01:46.580]<br />
entering different industries</p>
<p>[00:01:48.240]<br />
such as geothermal energy, environmental and civil.</p>
<p>[00:01:52.960]<br />
And in 2017, ARANZ Geo rebranded to Seequent.</p>
<p>[00:01:58.000]<br />
In 2018, Seequent acquired Geosoft</p>
<p>[00:02:00.670]<br />
the developers of Geophysical Software</p>
<p>[00:02:02.520]<br />
such as a Oasis montaj.</p>
<p>[00:02:04.520]<br />
And then in 2019, acquired GeoStudio,</p>
<p>[00:02:07.710]<br />
the developers of Geotechnical Analysis Software.</p>
<p>[00:02:11.930]<br />
During this time Seequent has continued developing</p>
<p>[00:02:15.630]<br />
our 3D geological modeling in Leapfrog</p>
<p>[00:02:18.620]<br />
and collaboration tools such as Central.</p>
<p>[00:02:22.480]<br />
Earlier this year,</p>
<p>[00:02:23.313]<br />
Seequent was acquired by Bentley Systems</p>
<p>[00:02:25.940]<br />
and has become a Bentley company</p>
<p>[00:02:27.433]<br />
which opens us up to even more possibilities</p>
<p>[00:02:30.430]<br />
for future development for geoscience solutions.</p>
<p>[00:02:36.310]<br />
Seequent have office globally</p>
<p>[00:02:37.990]<br />
with our head office in Christchurch, New Zealand.</p>
<p>[00:02:40.890]<br />
And more locally we have offices in Australia,</p>
<p>[00:02:43.550]<br />
in Perth and in Brisbane.</p>
<p>[00:02:48.240]<br />
Like I mentioned Seequent’s involved</p>
<p>[00:02:49.970]<br />
in number of different industries</p>
<p>[00:02:52.140]<br />
covering things from a contaminant modeling,</p>
<p>[00:02:55.550]<br />
road and toll construction,</p>
<p>[00:02:57.360]<br />
groundwater detection and management,</p>
<p>[00:02:59.460]<br />
geothermal exploration, resource evaluation</p>
<p>[00:03:02.560]<br />
and so much more.</p>
<p>[00:03:08.210]<br />
Today we’re looking at contaminant</p>
<p>[00:03:10.253]<br />
or contaminated and environmental projects.</p>
<p>[00:03:13.340]<br />
Seequent provides solutions across the life</p>
<p>[00:03:15.650]<br />
of contaminated site projects</p>
<p>[00:03:17.060]<br />
from the initial desk study phase</p>
<p>[00:03:19.140]<br />
to the site investigation</p>
<p>[00:03:20.800]<br />
through to the remediation, design and execution.</p>
<p>[00:03:25.070]<br />
These solutions include our geophysics software</p>
<p>[00:03:27.320]<br />
such as Oasis montaj and VOXI.</p>
<p>[00:03:29.930]<br />
Our 3D visualization and geological modeling</p>
<p>[00:03:32.490]<br />
and contaminant modeling in Leapfrog Works</p>
<p>[00:03:35.460]<br />
as well as flow and transport analysis in GeoStudio.</p>
<p>[00:03:39.710]<br />
Across all of this we have Seequent Central</p>
<p>[00:03:42.100]<br />
a cloud-based model management and collaboration tool</p>
<p>[00:03:45.320]<br />
for the life of the projects.</p>
<p>[00:03:48.090]<br />
Like I said, today we’re specifically looking</p>
<p>[00:03:50.220]<br />
at the Contaminants Extension in Leapfrog Works.</p>
<p>[00:03:56.040]<br />
So what is the Contaminants Extension?</p>
<p>[00:03:58.670]<br />
The Contaminants Extension</p>
<p>[00:03:59.790]<br />
is an optional module for Leapfrog works</p>
<p>[00:04:01.660]<br />
to enable spatial modeling of numeric data</p>
<p>[00:04:04.540]<br />
for defining the concentration, distribution</p>
<p>[00:04:07.330]<br />
and mass of the institute contamination at a site.</p>
<p>[00:04:11.910]<br />
The Seequent Contaminants Extension provides</p>
<p>[00:04:14.670]<br />
a transparent and auditable data-driven interpretation.</p>
<p>[00:04:19.900]<br />
It provides interactive visual tools</p>
<p>[00:04:22.430]<br />
with the extension that makes geostatistics</p>
<p>[00:04:24.760]<br />
accessible for all geoscientists</p>
<p>[00:04:27.610]<br />
ensuring that you and your teams</p>
<p>[00:04:29.120]<br />
can work the best with your data.</p>
<p>[00:04:32.600]<br />
It has built-in report tables</p>
<p>[00:04:34.730]<br />
and cross-section evaluations</p>
<p>[00:04:36.270]<br />
that are dynamically linked</p>
<p>[00:04:37.620]<br />
to your plume models and your data.</p>
<p>[00:04:41.350]<br />
So they will update automatically</p>
<p>[00:04:43.430]<br />
as the investigation and the project evolves in new day</p>
<p>[00:04:47.010]<br />
that comes into your Leapfrog project.</p>
<p>[00:04:50.320]<br />
It also means there’s an interrogatable estimation tool</p>
<p>[00:04:54.730]<br />
that provides easy access to the details</p>
<p>[00:04:58.260]<br />
for how you’ve created your informed decisions.</p>
<p>[00:05:02.595]<br />
In short, it allows you to take known data for your project</p>
<p>[00:05:07.070]<br />
and analyze it with spatial variability</p>
<p>[00:05:09.500]<br />
of the concentrations for your data</p>
<p>[00:05:12.390]<br />
and it helps you define your projects.</p>
<p>[00:05:20.860]<br />
So that’s what it is.</p>
<p>[00:05:22.410]<br />
Why did we develop it? And why is it useful?</p>
<p>[00:05:25.800]<br />
Well, we did it because we wanted to provide</p>
<p>[00:05:27.840]<br />
these accessible best practice solution tools</p>
<p>[00:05:30.690]<br />
for you to model your contamination plumes.</p>
<p>[00:05:34.290]<br />
We wanted to make characterizing contaminated land</p>
<p>[00:05:36.810]<br />
in groundwater, in a rigorous and auditable way</p>
<p>[00:05:39.730]<br />
that combines 3D dynamic geological Models</p>
<p>[00:05:42.700]<br />
with these best practice geostatistical methods.</p>
<p>[00:05:47.500]<br />
The ultimate goal of estimation</p>
<p>[00:05:48.970]<br />
is to combine the qualitative geological interpretation</p>
<p>[00:05:52.310]<br />
of your project.</p>
<p>[00:05:53.680]<br />
So your geological models</p>
<p>[00:05:55.270]<br />
with the sparsely sampled quantitative data</p>
<p>[00:05:58.450]<br />
to create spatial predictions</p>
<p>[00:06:00.130]<br />
for the distribution of that contaminant plan.</p>
<p>[00:06:09.140]<br />
The Contaminants Extension</p>
<p>[00:06:10.610]<br />
coupled with Seequent Central provides a wholistic way</p>
<p>[00:06:13.877]<br />
for the engineer and modeler for a project</p>
<p>[00:06:16.780]<br />
to effectively understand</p>
<p>[00:06:18.580]<br />
and importantly communicate the site conditions</p>
<p>[00:06:21.490]<br />
with both internal and external stakeholders</p>
<p>[00:06:24.310]<br />
in a collaborative environment.</p>
<p>[00:06:27.090]<br />
The graphic here shows an overview of the flow</p>
<p>[00:06:29.610]<br />
from Leapfrog to Central as part of the solution.</p>
<p>[00:06:34.350]<br />
Starting at the base here and moving up</p>
<p>[00:06:37.690]<br />
we can see that with your investigation data models</p>
<p>[00:06:40.982]<br />
and estimations can be created by the modeler</p>
<p>[00:06:44.260]<br />
and the environmental engineer for instance.</p>
<p>[00:06:48.310]<br />
And using Central,</p>
<p>[00:06:49.600]<br />
they can share this and collaborate on this</p>
<p>[00:06:53.000]<br />
with internal team members such as managers,</p>
<p>[00:06:55.830]<br />
reviewers or other modelers</p>
<p>[00:06:58.360]<br />
to build up the understanding of the site</p>
<p>[00:07:02.040]<br />
or as part of the project review process.</p>
<p>[00:07:06.420]<br />
At the very top of this infographic here</p>
<p>[00:07:08.260]<br />
we can then see how we can use Central</p>
<p>[00:07:10.330]<br />
to engage with external stakeholders</p>
<p>[00:07:13.130]<br />
such as the clients or other contractors</p>
<p>[00:07:16.650]<br />
and you can bring them into the Central project again</p>
<p>[00:07:18.740]<br />
to communicate in an interactive and visual way.</p>
<p>[00:07:24.820]<br />
The Seequent solution for the contaminant projects then</p>
<p>[00:07:27.490]<br />
is all about knowing the impact</p>
<p>[00:07:29.490]<br />
of environmental contamination for your projects.</p>
<p>[00:07:32.880]<br />
Through first seeing the problem</p>
<p>[00:07:34.360]<br />
in a clear, intuitive and visual way.</p>
<p>[00:07:38.030]<br />
The picture is worth 1,000 words</p>
<p>[00:07:39.440]<br />
a 3D visualization of the site has to be worth a million.</p>
<p>[00:07:44.260]<br />
With a Contaminants Extension,</p>
<p>[00:07:45.810]<br />
it’s an interactive dynamic site model</p>
<p>[00:07:47.990]<br />
you can create and understand</p>
<p>[00:07:50.130]<br />
and aid in your analysis and your assumptions</p>
<p>[00:07:53.070]<br />
and recommendations for your reports.</p>
<p>[00:08:01.340]<br />
I’ll now hand over to Steve</p>
<p>[00:08:03.500]<br />
who will take us through Leapfrog</p>
<p>[00:08:05.540]<br />
and the Contaminants Extension.</p>
<p>[00:08:27.267]<br />
So we’ll just switch over from the slides here. (mumbles).</p>
<p>[00:08:38.840]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Thanks very much Aaron<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:08:40.756]<br />
for that great introduction.</p>
<p>[00:08:42.290]<br />
So I’m going to go through the</p>
<p>[00:08:44.470]<br />
Contaminants Extension Workflow.</p>
<p>[00:08:48.830]<br />
The end result is a resource model</p>
<p>[00:08:50.820]<br />
and through the whole process we have a look at our data.</p>
<p>[00:08:53.770]<br />
We go through the mining,</p>
<p>[00:08:55.340]<br />
run through estimations and validate and report at the end.</p>
<p>[00:08:59.650]<br />
Throughout the whole process it’s not a linear</p>
<p>[00:09:01.960]<br />
it can be a cyclical process</p>
<p>[00:09:03.840]<br />
so that whilst we’re at any stage</p>
<p>[00:09:06.310]<br />
we can always go back to our geology</p>
<p>[00:09:08.480]<br />
and look at how things are relating to each other.</p>
<p>[00:09:15.580]<br />
We’re going to go through a couple of key steps</p>
<p>[00:09:18.950]<br />
that we need to consider whilst building up a plume model.</p>
<p>[00:09:24.070]<br />
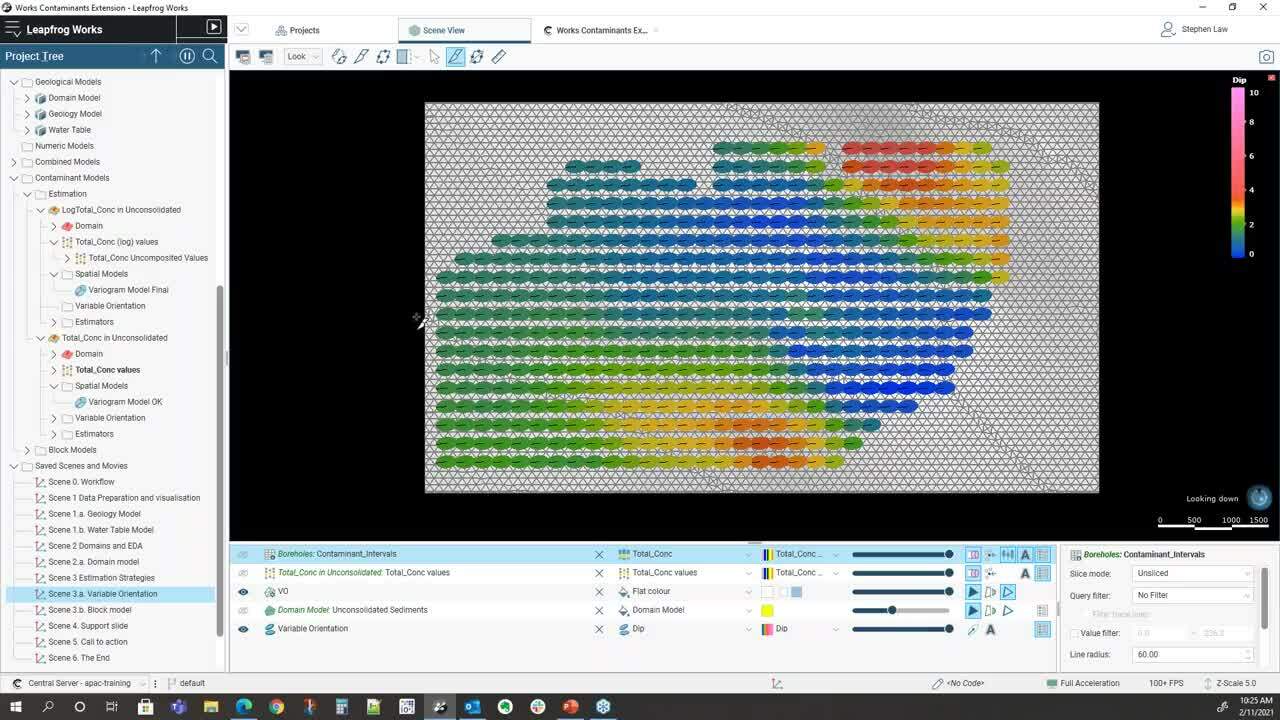
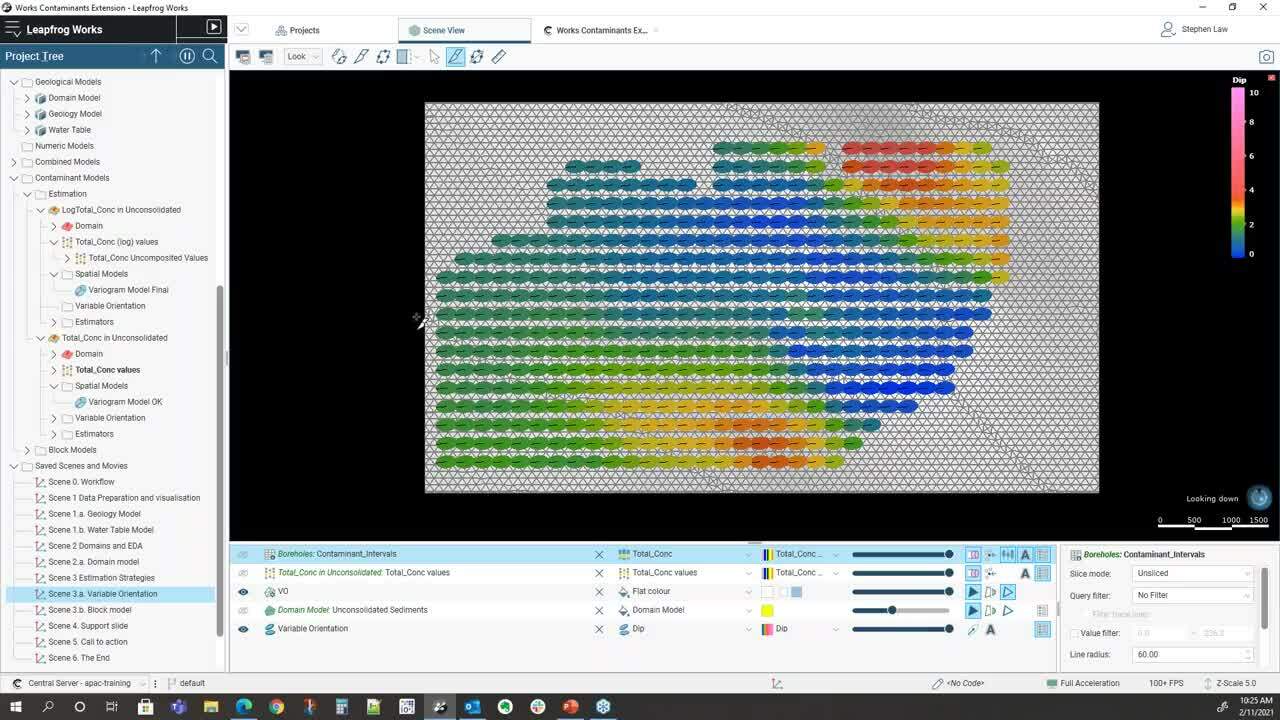
So the first part is being well aware of our data</p>
<p>[00:09:27.960]<br />
and how we can visualize it very easily</p>
<p>[00:09:30.240]<br />
within Leapfrog Works.</p>
<p>[00:09:31.980]<br />
So we want to consider things,</p>
<p>[00:09:33.659]<br />
where is our data and how is it located especially?</p>
<p>[00:09:37.500]<br />
We have tools in-built that we can check</p>
<p>[00:09:39.640]<br />
and make sure our data is validated</p>
<p>[00:09:42.490]<br />
and we have really good visualization tools.</p>
<p>[00:09:47.360]<br />
So within this project we have three models,</p>
<p>[00:09:51.440]<br />
the first one is a geology model</p>
<p>[00:09:53.840]<br />
and you can see here that it’s got outwash</p>
<p>[00:09:59.580]<br />
and ice deposits until above a bedrock sequence.</p>
<p>[00:10:03.960]<br />
And within that we have a range of vertical boreholes</p>
<p>[00:10:07.210]<br />
and these have been sampled generally in isolated locations</p>
<p>[00:10:13.840]<br />
for a chlorine contaminant</p>
<p>[00:10:16.030]<br />
and that’s what we’re going to be evaluating.</p>
<p>[00:10:19.240]<br />
As well as the geology model,</p>
<p>[00:10:21.440]<br />
we have a water table model.</p>
<p>[00:10:24.870]<br />
And this has been divided into a saturated</p>
<p>[00:10:28.040]<br />
and made (mumbles) again above bedrock.</p>
<p>[00:10:31.740]<br />
So one of the first things we want to consider is,</p>
<p>[00:10:34.910]<br />
is there any specific way that we should domain our data?</p>
<p>[00:10:38.040]<br />
To ask some of them, is chlorine associated with pathology?</p>
<p>[00:10:42.960]<br />
Is it associated with water table or a combination of both?</p>
<p>[00:10:47.199]<br />
And we have some tools within Leapfrog Works</p>
<p>[00:10:49.460]<br />
that can show this.</p>
<p>[00:10:52.180]<br />
So up in the boreholes’ database here,</p>
<p>[00:10:55.010]<br />
I have a table with the contaminants</p>
<p>[00:11:00.320]<br />
and then I’ve got a geology table.</p>
<p>[00:11:02.370]<br />
We have a merge table function</p>
<p>[00:11:04.090]<br />
so I’ve been able to merge these together.</p>
<p>[00:11:06.370]<br />
And this enables me then to evaluate them</p>
<p>[00:11:10.700]<br />
using some statistical tools</p>
<p>[00:11:13.120]<br />
in this case I’m using a box plot.</p>
<p>[00:11:17.640]<br />
And so this was here as a box plot of the geology</p>
<p>[00:11:20.360]<br />
against the contaminant.</p>
<p>[00:11:22.610]<br />
And we can see here that that is telling me</p>
<p>[00:11:24.910]<br />
that the outwash and local ice to contact deposits</p>
<p>[00:11:28.310]<br />
and the two, have the highest concentration of chlorine.</p>
<p>[00:11:32.310]<br />
And then there’s a lot less amount</p>
<p>[00:11:34.350]<br />
in the outwash deposits and bedrock.</p>
<p>[00:11:38.830]<br />
We can also have a look at this as a table of statistics.</p>
<p>[00:11:48.536]<br />
Excuse me.</p>
<p>[00:11:58.460]<br />
An idea of…</p>
<p>[00:11:59.810]<br />
We have a little bit total concentration.</p>
<p>[00:12:02.200]<br />
I can see the actual values within each one.</p>
<p>[00:12:06.220]<br />
One of the key things here is</p>
<p>[00:12:07.510]<br />
looking at the number of samples</p>
<p>[00:12:09.010]<br />
so we have only two samples within the bedrock.</p>
<p>[00:12:12.060]<br />
We’d identified that the outwash and ice deposits</p>
<p>[00:12:14.920]<br />
and the two had the greatest concentrations</p>
<p>[00:12:20.080]<br />
and we’ve got the most number of samples in those as well</p>
<p>[00:12:22.650]<br />
so we’ve got approximately 70 samples within the ice.</p>
<p>[00:12:27.310]<br />
And we’ve got the mains</p>
<p>[00:12:28.780]<br />
so we’ve got much higher means within ice</p>
<p>[00:12:32.660]<br />
and very low within the outwash deposits</p>
<p>[00:12:35.760]<br />
and again quite low</p>
<p>[00:12:36.970]<br />
but there’s only two samples within bedrock.</p>
<p>[00:12:41.280]<br />
So in this instance there’s no clear</p>
<p>[00:12:46.220]<br />
if we have a look at (mumbles)</p>
<p>[00:12:49.820]<br />
water table, we can see that both are saturated</p>
<p>[00:12:54.180]<br />
and the vital sign has significant concentrations</p>
<p>[00:12:59.030]<br />
and there’s not one particularly higher than the other.</p>
<p>[00:13:01.610]<br />
Saturated is a little bit more,</p>
<p>[00:13:03.600]<br />
but it’s not obviously completely different.</p>
<p>[00:13:08.900]<br />
So the way we’ll consider this then</p>
<p>[00:13:10.870]<br />
is I’m just going to separate a single domain</p>
<p>[00:13:13.940]<br />
that’s above bedrock and not taking into account</p>
<p>[00:13:16.790]<br />
whether it’s saturated or (mumbles).</p>
<p>[00:13:20.030]<br />
In this way, I have a geological model then</p>
<p>[00:13:23.370]<br />
that is just domained.</p>
<p>[00:13:37.140]<br />
So, once we’ve decided on</p>
<p>[00:13:38.630]<br />
how we’re going to domain our data,</p>
<p>[00:13:40.620]<br />
then we start using the Contaminants Extension information</p>
<p>[00:13:43.990]<br />
and we can link it to our geology model</p>
<p>[00:13:46.040]<br />
right through to the resource estimate.</p>
<p>[00:13:48.060]<br />
And we can do exploratory data analysis</p>
<p>[00:13:50.400]<br />
specifically on the domain itself.</p>
<p>[00:13:53.635]<br />
So we have a look at the domain model.</p>
<p>[00:13:57.190]<br />
And in this case</p>
<p>[00:13:58.023]<br />
this has now been all of the mythologies about bedrock</p>
<p>[00:14:02.320]<br />
have been combined into a single domain</p>
<p>[00:14:03.660]<br />
which is the yellow one on the scene.</p>
<p>[00:14:06.370]<br />
And we can see that the majority of the samples</p>
<p>[00:14:08.147]<br />
are within that and there’s just the one isolated to</p>
<p>[00:14:12.140]<br />
below in the bedrock which we won’t be considering</p>
<p>[00:14:14.960]<br />
in this particular estimate.</p>
<p>[00:14:24.890]<br />
When we build up estimation,</p>
<p>[00:14:29.130]<br />
we have what’s termed as contaminants model folder.</p>
<p>[00:14:32.230]<br />
So when we activate the Contaminants Extension License,</p>
<p>[00:14:37.600]<br />
it displays up as a folder in your project tree</p>
<p>[00:14:41.380]<br />
and there’s a estimation sub folder</p>
<p>[00:14:43.347]<br />
and a block model’s folder.</p>
<p>[00:14:45.490]<br />
The estimation folder can be considered to be</p>
<p>[00:14:47.950]<br />
where you set up all your parameters.</p>
<p>[00:14:49.930]<br />
And then the block model is where we display</p>
<p>[00:14:52.500]<br />
all of our results</p>
<p>[00:14:53.440]<br />
and we also do all our reporting and validation</p>
<p>[00:14:56.140]<br />
from that point as well.</p>
<p>[00:14:58.520]<br />
So I’ve decided to do a title constraint traction</p>
<p>[00:15:02.410]<br />
of this chloride within the unconsolidated domain</p>
<p>[00:15:05.870]<br />
which is this yellow block here.</p>
<p>[00:15:08.010]<br />
So when we create a domain estimator,</p>
<p>[00:15:12.920]<br />
we just select any closed solid within our project</p>
<p>[00:15:17.040]<br />
and you can see here that</p>
<p>[00:15:17.890]<br />
this is the unconsolidated sediments</p>
<p>[00:15:21.430]<br />
within our domain model.</p>
<p>[00:15:24.160]<br />
And then the numeric values are coming directly</p>
<p>[00:15:26.940]<br />
from our essay table, contaminant intervals,</p>
<p>[00:15:30.681]<br />
total concentration.</p>
<p>[00:15:32.760]<br />
So we can choose any numeric column</p>
<p>[00:15:34.860]<br />
within any part of our database.</p>
<p>[00:15:36.860]<br />
And also if we were storing point data</p>
<p>[00:15:39.520]<br />
within the points folder over here,</p>
<p>[00:15:41.630]<br />
we would be able to access that as well.</p>
<p>[00:15:43.980]<br />
So it doesn’t have to be within drew holes .</p>
<p>[00:15:48.720]<br />
I’m not applying any transform at this stage</p>
<p>[00:15:51.680]<br />
and we also have the capability of doing compositing.</p>
<p>[00:15:55.340]<br />
This data set doesn’t render itself to compositing</p>
<p>[00:15:58.790]<br />
because it’s got isolated values within each holes</p>
<p>[00:16:02.640]<br />
really only got one or two samples</p>
<p>[00:16:05.628]<br />
compositing small for if we’ve got a</p>
<p>[00:16:08.080]<br />
continuous range of samples down the borehole,</p>
<p>[00:16:12.020]<br />
and we’d want to composite that into large intervals</p>
<p>[00:16:17.510]<br />
to reduce their variance.</p>
<p>[00:16:20.170]<br />
So within each estimation for each domain,</p>
<p>[00:16:22.730]<br />
we have this sub folder.</p>
<p>[00:16:24.520]<br />
And from there we can see the domain itself.</p>
<p>[00:16:28.280]<br />
So we can double check that that is the volume of interest</p>
<p>[00:16:33.030]<br />
and we can display the values within that domain.</p>
<p>[00:16:37.700]<br />
So they have become point values at this stage</p>
<p>[00:16:42.800]<br />
and you can label those</p>
<p>[00:16:44.820]<br />
and you can also do statistics on these.</p>
<p>[00:16:47.610]<br />
So these now become domain statistics.</p>
<p>[00:16:50.500]<br />
So by right clicking on the values,</p>
<p>[00:16:54.280]<br />
I get a histogram of those values there.</p>
<p>[00:16:58.910]<br />
As with all Leapfrog Works graphs,</p>
<p>[00:17:04.010]<br />
they are interacting with the 3D scene.</p>
<p>[00:17:07.130]<br />
So let’s just move this over here.</p>
<p>[00:17:12.920]<br />
If I wanted to find out where these core values were,</p>
<p>[00:17:17.590]<br />
I can just highlight the one that on the graph</p>
<p>[00:17:20.610]<br />
and that will show up in the same format.</p>
<p>[00:17:23.450]<br />
Let’s do these ones about 100, there they are there.</p>
<p>[00:17:27.350]<br />
So gives us then idea of whether or not</p>
<p>[00:17:30.160]<br />
we need to create sub domains.</p>
<p>[00:17:32.400]<br />
If all the high grades were focused a little bit more</p>
<p>[00:17:35.520]<br />
we might be able to subdivide them out.</p>
<p>[00:17:37.210]<br />
But in this case,</p>
<p>[00:17:38.630]<br />
we’re just going to treat everything in the one domain.</p>
<p>[00:17:44.870]<br />
The special models folder is where we can do variography</p>
<p>[00:17:49.990]<br />
which is a spatial analysis of the data</p>
<p>[00:17:52.670]<br />
to see whether there are any specific directions</p>
<p>[00:17:55.320]<br />
where the concentration grades might be aligned.</p>
<p>[00:18:02.980]<br />
I’ll open this one up.</p>
<p>[00:18:12.510]<br />
So the way the variography works</p>
<p>[00:18:14.320]<br />
is that we select the direction</p>
<p>[00:18:16.700]<br />
and then we model it in three directions.</p>
<p>[00:18:21.370]<br />
One with the greatest degree of continuity et cetera.</p>
<p>[00:18:25.160]<br />
This radio plot is a plane within this.</p>
<p>[00:18:29.390]<br />
We’re looking in a fairly horizontal plane here</p>
<p>[00:18:32.667]<br />
and you can see that we have…</p>
<p>[00:18:35.600]<br />
we can always see the ellipsoid that we’re working with</p>
<p>[00:18:38.900]<br />
so to see whether it’s in the like</p>
<p>[00:18:41.160]<br />
we wanted to make that a little bit flatter.</p>
<p>[00:18:43.250]<br />
We didn’t leave it in a 3D scene</p>
<p>[00:18:45.660]<br />
and it will update on the form here</p>
<p>[00:18:48.210]<br />
and the graphs will update.</p>
<p>[00:18:51.260]<br />
We can use this radial plot.</p>
<p>[00:18:54.520]<br />
This may suggest that this orientation</p>
<p>[00:18:57.210]<br />
has a little bit more continuity and we’ll re-adjust.</p>
<p>[00:19:02.520]<br />
This particular data set doesn’t create</p>
<p>[00:19:05.680]<br />
a very good experimental variogram</p>
<p>[00:19:08.130]<br />
so we do have some other options</p>
<p>[00:19:09.710]<br />
where we could use a log transform prior to that</p>
<p>[00:19:14.293]<br />
so here I’ve done a log transform on the values</p>
<p>[00:19:19.450]<br />
and that was simply done by at the very beginning.</p>
<p>[00:19:22.550]<br />
When I selected my samples</p>
<p>[00:19:24.256]<br />
I could apply a log transform in through here.</p>
<p>[00:19:28.630]<br />
This gets rid of some of the variability</p>
<p>[00:19:30.650]<br />
especially the range and extreme differences</p>
<p>[00:19:34.350]<br />
between very high values and very low values</p>
<p>[00:19:37.260]<br />
and this can help develop a better semi-variably model</p>
<p>[00:19:42.300]<br />
if we have a little bit this one.</p>
<p>[00:19:50.624]<br />
You can see here that the semivariogram</p>
<p>[00:19:53.240]<br />
is a little bit better structured than it was before.</p>
<p>[00:19:56.700]<br />
Again this data is quite sparse.</p>
<p>[00:19:58.760]<br />
So this is about the best we could do</p>
<p>[00:20:01.070]<br />
with this particular data set.</p>
<p>[00:20:04.090]<br />
We have the ability to pick a couple of range of structures,</p>
<p>[00:20:08.970]<br />
we have a nugget plus a potentially two structures</p>
<p>[00:20:12.370]<br />
but in this case I’ll just pick a nugget,</p>
<p>[00:20:15.360]<br />
a fairly large nugget and a single structure</p>
<p>[00:20:18.030]<br />
which is sufficient to model this.</p>
<p>[00:20:21.190]<br />
We can get an idea of that model.</p>
<p>[00:20:23.350]<br />
So this is in the…</p>
<p>[00:20:25.090]<br />
The range is quite large, 1200 to 1400 meters.</p>
<p>[00:20:29.920]<br />
And that’s because the data itself is quite widely spaced.</p>
<p>[00:20:33.620]<br />
And we’ve only got a very short range</p>
<p>[00:20:35.390]<br />
in the Z direction because we’re looking at a</p>
<p>[00:20:37.294]<br />
fairly constrained tabular body of potential contamination.</p>
<p>[00:20:44.340]<br />
The advantage to building the variography</p>
<p>[00:20:46.390]<br />
within the contaminants extension is that</p>
<p>[00:20:49.070]<br />
everything is self-contained,</p>
<p>[00:20:50.410]<br />
and once we hit the side button,</p>
<p>[00:20:52.830]<br />
then that variogram is all ready to use</p>
<p>[00:20:56.510]<br />
within the estimations itself.</p>
<p>[00:21:00.480]<br />
We also have a concept of variable orientation.</p>
<p>[00:21:04.750]<br />
Sometimes the design of the Contaminant</p>
<p>[00:21:10.520]<br />
isn’t purely tabular</p>
<p>[00:21:12.480]<br />
and there may be local irregularities</p>
<p>[00:21:14.810]<br />
within the surfaces.</p>
<p>[00:21:16.680]<br />
So we can use those surfaces</p>
<p>[00:21:18.450]<br />
to help guide the local orientation of the variogram</p>
<p>[00:21:22.750]<br />
when we’re doing sample selection.</p>
<p>[00:21:25.470]<br />
So just don’t want it here already.</p>
<p>[00:21:31.890]<br />
So the way we do that</p>
<p>[00:21:32.810]<br />
is that we set up a reference surface</p>
<p>[00:21:35.440]<br />
and in this case it’s the actual base</p>
<p>[00:21:37.570]<br />
over the top of the bedrock</p>
<p>[00:21:39.530]<br />
and we can see here that we’ve colored this by a dip.</p>
<p>[00:21:44.480]<br />
So we can see that we’ve got some varying dips</p>
<p>[00:21:47.670]<br />
that flat up in the middle</p>
<p>[00:21:49.070]<br />
and slightly more dips around the edges.</p>
<p>[00:21:51.830]<br />
If I can hop and look cross sections through there,</p>
<p>[00:22:03.450]<br />
we can see that what will happen</p>
<p>[00:22:06.610]<br />
is when we pick out samples,</p>
<p>[00:22:09.060]<br />
the ellipsoid will be orientated as local</p>
<p>[00:22:12.780]<br />
and so overall ellipsoid is in this general direction,</p>
<p>[00:22:16.010]<br />
but where it gets flat out,</p>
<p>[00:22:17.400]<br />
it will rotate in a flat orientation.</p>
<p>[00:22:21.570]<br />
The actual ratios and the variogram parameters themselves</p>
<p>[00:22:24.960]<br />
are still maintained.</p>
<p>[00:22:26.267]<br />
It’s just the orientation</p>
<p>[00:22:28.260]<br />
for when the samples are being selected.</p>
<p>[00:22:35.270]<br />
When we’re doing an estimation,</p>
<p>[00:22:36.730]<br />
we have to consider a few different things.</p>
<p>[00:22:39.330]<br />
So we need to understand what the stage our project is</p>
<p>[00:22:42.650]<br />
and this might help us work out</p>
<p>[00:22:44.520]<br />
which estimation method to use.</p>
<p>[00:22:46.550]<br />
If we’ve got early stage,</p>
<p>[00:22:48.190]<br />
we might not have very much data.</p>
<p>[00:22:49.910]<br />
We might not have sufficient data to get a variogram.</p>
<p>[00:22:52.740]<br />
So we will need to use either nearest neighbor</p>
<p>[00:22:55.510]<br />
or inverse distance type methods.</p>
<p>[00:22:59.476]<br />
There can be a lot of work to go into the search strategy.</p>
<p>[00:23:02.280]<br />
And that is how many samples are we going to use</p>
<p>[00:23:05.061]<br />
to define a local estimate.</p>
<p>[00:23:08.850]<br />
And then once we run our estimate,</p>
<p>[00:23:10.900]<br />
we need to validate it against the original data</p>
<p>[00:23:14.250]<br />
to make sure that we’re maintaining the same kind of means</p>
<p>[00:23:17.950]<br />
and then we want to report our results at the end.</p>
<p>[00:23:23.150]<br />
To set up the estimators,</p>
<p>[00:23:25.970]<br />
we work through this estimators sub folder here</p>
<p>[00:23:29.760]<br />
and if you right click</p>
<p>[00:23:30.920]<br />
you can go have inverse distance nearest neighbor kriging</p>
<p>[00:23:34.857]<br />
and we can do ordinary kriging, simple kriging</p>
<p>[00:23:38.010]<br />
or we can use the actual RBF estimator as well.</p>
<p>[00:23:41.520]<br />
Which can be quite good</p>
<p>[00:23:42.870]<br />
for when we haven’t got sufficient data to a variogram.</p>
<p>[00:23:47.110]<br />
It’s a global estimator.</p>
<p>[00:23:49.540]<br />
Using the same underlying algorithm</p>
<p>[00:23:51.470]<br />
that is used for when we krig the surfaces</p>
<p>[00:23:54.170]<br />
or the numeric models that can be run</p>
<p>[00:23:56.600]<br />
within Leapfrog Works itself.</p>
<p>[00:24:00.710]<br />
To set one up</p>
<p>[00:24:01.990]<br />
once I open we can edit that,</p>
<p>[00:24:04.650]<br />
we can apply clipping which is</p>
<p>[00:24:08.390]<br />
if we want to constrain the very high values</p>
<p>[00:24:11.010]<br />
and give them less influence,</p>
<p>[00:24:12.550]<br />
we can apply that on floor.</p>
<p>[00:24:15.090]<br />
In this case because I’ve actually looking at contaminants</p>
<p>[00:24:17.380]<br />
I don’t really want to cut them down initially</p>
<p>[00:24:19.580]<br />
so I will leave them unclicked.</p>
<p>[00:24:22.540]<br />
I’m just doing point kriging in this case</p>
<p>[00:24:24.710]<br />
so just I’m not using discretization at all.</p>
<p>[00:24:28.590]<br />
In some cases if I want to block kriging,</p>
<p>[00:24:30.460]<br />
then I would increase these discretization amounts.</p>
<p>[00:24:35.170]<br />
I’m using ordinary kriging.</p>
<p>[00:24:36.693]<br />
This is where we determine ordinary or simple.</p>
<p>[00:24:41.120]<br />
And this is the variogram.</p>
<p>[00:24:42.750]<br />
So you can pick.</p>
<p>[00:24:44.180]<br />
It will see the variogram within the folder.</p>
<p>[00:24:50.460]<br />
You set to a particular variogram</p>
<p>[00:24:53.090]<br />
and that’s more for making sure</p>
<p>[00:24:55.730]<br />
that it’s set to the correct orientation.</p>
<p>[00:24:58.240]<br />
And then it’ll automatically go to the maximum range</p>
<p>[00:25:01.100]<br />
of that variogram but you can modify these</p>
<p>[00:25:03.680]<br />
to be less than that or greater.</p>
<p>[00:25:05.790]<br />
Depends on the circumstances</p>
<p>[00:25:08.750]<br />
and often you’ll do a lot of testing</p>
<p>[00:25:11.230]<br />
and sensitivity analysis to determine</p>
<p>[00:25:13.760]<br />
the best scope for this one.</p>
<p>[00:25:18.400]<br />
And then we can choose</p>
<p>[00:25:19.890]<br />
minimum and maximum number of samples.</p>
<p>[00:25:22.610]<br />
And this effectively becomes</p>
<p>[00:25:24.280]<br />
our grade variable within the block model</p>
<p>[00:25:27.710]<br />
and we can store other variables as well.</p>
<p>[00:25:30.510]<br />
So if we’re using kriging,</p>
<p>[00:25:32.030]<br />
we can store things such as kriging variance,</p>
<p>[00:25:34.240]<br />
slope and regression</p>
<p>[00:25:35.820]<br />
which would go as to how good the estimate might be.</p>
<p>[00:25:39.430]<br />
We can store things such as average distance to samples</p>
<p>[00:25:42.290]<br />
and these may help us with classification</p>
<p>[00:25:44.810]<br />
criteria down the track.</p>
<p>[00:25:47.150]<br />
So I’ve created us an ordinary kriged estimator here</p>
<p>[00:25:52.640]<br />
and then I’ve also got one.</p>
<p>[00:25:54.270]<br />
This one is exactly the same,</p>
<p>[00:25:57.340]<br />
but in this case I’ve actually applied</p>
<p>[00:25:59.610]<br />
the variable orientation.</p>
<p>[00:26:01.230]<br />
So we’ve set up a variable orientation within the folder.</p>
<p>[00:26:04.190]<br />
It will be accessible in our parameter set up through here</p>
<p>[00:26:09.050]<br />
and I’ll give it a slightly different name.</p>
<p>[00:26:11.550]<br />
This one I will need to compare the kriging results</p>
<p>[00:26:17.310]<br />
with and without variable orientation apply.</p>
<p>[00:26:21.570]<br />
So we set up all of our different parameters</p>
<p>[00:26:24.250]<br />
in this estimation folder.</p>
<p>[00:26:27.360]<br />
And we are then able to create a block model</p>
<p>[00:26:32.690]<br />
to look at the results.</p>
<p>[00:26:40.300]<br />
Setting up a block model is quite simple</p>
<p>[00:26:43.440]<br />
so we can create a new block model</p>
<p>[00:26:45.900]<br />
and we can also import block models from other software.</p>
<p>[00:26:49.790]<br />
And when we’ve created one,</p>
<p>[00:26:51.990]<br />
we can edit it by opening it up.</p>
<p>[00:26:55.500]<br />
And in this case,</p>
<p>[00:26:56.333]<br />
we’ve got a block size of 80 by 80 meters</p>
<p>[00:26:58.720]<br />
in the X, Y direction and five in the Z</p>
<p>[00:27:02.260]<br />
so we’re looking at that fairly thin.</p>
<p>[00:27:04.270]<br />
And if we’re looking at it again very visual</p>
<p>[00:27:07.167]<br />
and we can see our block size.</p>
<p>[00:27:13.230]<br />
So this is 80 by 80 in the X Y direction here.</p>
<p>[00:27:18.400]<br />
And we can change these boundaries at any time.</p>
<p>[00:27:20.810]<br />
So once it’s been made,</p>
<p>[00:27:21.900]<br />
if we wanted to adjust it,</p>
<p>[00:27:23.533]<br />
I could just move the boundary up</p>
<p>[00:27:25.720]<br />
and it will automatically update the model.</p>
<p>[00:27:28.930]<br />
I want to do that now so it doesn’t rerun.</p>
<p>[00:27:31.820]<br />
A key concept when working with block models</p>
<p>[00:27:34.030]<br />
within the Contaminants Extension is this evaluations tab.</p>
<p>[00:27:38.260]<br />
So basically whatever we want to flag into the block model</p>
<p>[00:27:41.794]<br />
needs to come across from the left-hand side</p>
<p>[00:27:44.590]<br />
across to the right.</p>
<p>[00:27:46.040]<br />
So what I’m doing here</p>
<p>[00:27:47.090]<br />
is I’m flagging each block centroid with the domain model</p>
<p>[00:27:50.980]<br />
so that will flag it with whether it’s within that domain</p>
<p>[00:27:56.590]<br />
but we can also flag at geology.</p>
<p>[00:27:58.410]<br />
So this will put into whether it’s two</p>
<p>[00:28:00.770]<br />
or whether it’s bedrock</p>
<p>[00:28:02.610]<br />
and also avoid us saturated sides</p>
<p>[00:28:05.110]<br />
could be flagged as well.</p>
<p>[00:28:07.500]<br />
And then this is Y grade or contaminant variables here.</p>
<p>[00:28:12.870]<br />
So I’m just putting in all four that are generated</p>
<p>[00:28:17.200]<br />
so I’ve got an inverse distance,</p>
<p>[00:28:19.360]<br />
I’ve got two different krigings.</p>
<p>[00:28:21.734]<br />
And also the nearest neighbor.</p>
<p>[00:28:23.460]<br />
Nearest neighbor can be useful</p>
<p>[00:28:25.140]<br />
for helping with validation of the model.</p>
<p>[00:28:29.480]<br />
Again, any of these can be removed or added at one time.</p>
<p>[00:28:32.680]<br />
So if I did want to look at the RBF,</p>
<p>[00:28:35.070]<br />
I would just bring it across</p>
<p>[00:28:37.170]<br />
and then the model would rerun.</p>
<p>[00:28:41.200]<br />
So we can see here, the model has been generated.</p>
<p>[00:28:45.710]<br />
And one of the key things for validation of first</p>
<p>[00:28:49.120]<br />
is to actually look at the samples against the results.</p>
<p>[00:28:54.550]<br />
So if I just go into a cross section</p>
<p>[00:29:06.010]<br />
making sure that our color schemes</p>
<p>[00:29:08.760]<br />
forbides the data and the bot model are the same</p>
<p>[00:29:11.940]<br />
and we can import these colors</p>
<p>[00:29:13.580]<br />
so if we’ve got colors defined for our essays up here,</p>
<p>[00:29:23.952]<br />
we can export those colors</p>
<p>[00:29:25.640]<br />
like create a little file,</p>
<p>[00:29:27.360]<br />
and then we can go down to our block model</p>
<p>[00:29:29.660]<br />
and we can import those colors down here.</p>
<p>[00:29:34.420]<br />
So I did on this one, look at that colors import</p>
<p>[00:29:38.370]<br />
and then pick up that file.</p>
<p>[00:29:39.670]<br />
And that way we get exactly same color scheme.</p>
<p>[00:29:42.640]<br />
And what we’re hoping to see</p>
<p>[00:29:43.910]<br />
is a little we’ve got yellows</p>
<p>[00:29:46.610]<br />
we should hopefully have orange or yellows around,</p>
<p>[00:29:49.710]<br />
higher grades will be in the reds and purples</p>
<p>[00:29:53.610]<br />
and lower grades should show up.</p>
<p>[00:30:02.308]<br />
Of course we have a lot of data in here.</p>
<p>[00:30:06.074]<br />
There we go.</p>
<p>[00:30:06.907]<br />
So we can see some low grade points over here</p>
<p>[00:30:09.280]<br />
and that’s equating.</p>
<p>[00:30:11.170]<br />
We’ve got really good tools for looking at the details</p>
<p>[00:30:14.870]<br />
of the results on a block-by-block basis.</p>
<p>[00:30:17.720]<br />
If I highlighted this particular block here,</p>
<p>[00:30:21.250]<br />
we have this interrogate estimator function.</p>
<p>[00:30:26.130]<br />
And this is the estimator that it’s looking at.</p>
<p>[00:30:29.080]<br />
So like ordinary krig without variable orientation</p>
<p>[00:30:32.560]<br />
it’s found there’s many samples</p>
<p>[00:30:35.390]<br />
and depending on how we’ve set up our search,</p>
<p>[00:30:38.120]<br />
in this case it’s included all those samples.</p>
<p>[00:30:41.050]<br />
It’s showing us the kriging lights</p>
<p>[00:30:42.360]<br />
that have been applied and we can sort these</p>
<p>[00:30:45.177]<br />
and distances from the sample.</p>
<p>[00:30:48.660]<br />
So we can see the closest sample used was 175 meters away</p>
<p>[00:30:53.080]<br />
and the greatest one was over 1,000 meters away.</p>
<p>[00:30:56.200]<br />
But it’s got lights through there.</p>
<p>[00:31:00.470]<br />
Part of the kriging process</p>
<p>[00:31:01.730]<br />
can give negative kriging weights</p>
<p>[00:31:03.670]<br />
and that is not a problem</p>
<p>[00:31:04.850]<br />
unless we start to see negative grade values.</p>
<p>[00:31:07.770]<br />
In that case, you might need to change</p>
<p>[00:31:09.520]<br />
our sample selection a little bit</p>
<p>[00:31:12.660]<br />
to eradicate that influence.</p>
<p>[00:31:16.420]<br />
That’s just part of the kriging process.</p>
<p>[00:31:21.130]<br />
You can see also that when we do that we can…</p>
<p>[00:31:23.740]<br />
it shows the ellipsoid for that area.</p>
<p>[00:31:26.720]<br />
And if we filter the block model</p>
<p>[00:31:30.818]<br />
and I’ll just take away the ellipsoid for a second,</p>
<p>[00:31:36.180]<br />
it shows you the block and it shows you the samples</p>
<p>[00:31:39.113]<br />
that are being used.</p>
<p>[00:31:40.540]<br />
So you’ve got a nice spatial visualization</p>
<p>[00:31:44.000]<br />
of what samples are actually being used</p>
<p>[00:31:46.080]<br />
to estimate the grade within that box through there.</p>
<p>[00:31:48.947]<br />
And if you want to try something else,</p>
<p>[00:31:50.760]<br />
you can go directly to the estimator and modify.</p>
<p>[00:31:55.360]<br />
If I want to change this to three samples,</p>
<p>[00:31:58.070]<br />
reduce these chains to search</p>
<p>[00:32:00.600]<br />
and you can get see what the impacts could be. Okay.</p>
<p>[00:32:06.790]<br />
So apart from visualization</p>
<p>[00:32:08.910]<br />
and looking at on a block-by-block basis,</p>
<p>[00:32:12.620]<br />
we can also run swath plots.</p>
<p>[00:32:18.780]<br />
So within the block model itself,</p>
<p>[00:32:20.870]<br />
we can create a new swath plot.</p>
<p>[00:32:23.210]<br />
And the beauty of this is that</p>
<p>[00:32:25.200]<br />
they are embedded within the block model</p>
<p>[00:32:27.090]<br />
so that when we update that data later on,</p>
<p>[00:32:29.750]<br />
the swath plots will update as well</p>
<p>[00:32:32.270]<br />
and the same goes for reports.</p>
<p>[00:32:34.060]<br />
If we build a report on the model</p>
<p>[00:32:36.170]<br />
then if we change the data,</p>
<p>[00:32:37.650]<br />
then the reports will have tables.</p>
<p>[00:32:40.409]<br />
So we’ll have a look at a swath</p>
<p>[00:32:42.610]<br />
that’s already been prepared</p>
<p>[00:32:44.030]<br />
and you can store any number of swaths within the model.</p>
<p>[00:32:49.330]<br />
So what this one is doing</p>
<p>[00:32:50.530]<br />
is it’s looking at the inverse distance.</p>
<p>[00:32:53.470]<br />
It’s got the nearest neighbor which is this gray</p>
<p>[00:32:56.170]<br />
and then we’ve got two different types of kriging.</p>
<p>[00:32:59.950]<br />
So we can see straight away.</p>
<p>[00:33:01.320]<br />
And it’s showing us in the X, Y and Z directions</p>
<p>[00:33:05.810]<br />
relative to the axis of the block model.</p>
<p>[00:33:08.830]<br />
So if we look in the X direction here,</p>
<p>[00:33:10.640]<br />
we can see that all three estimators</p>
<p>[00:33:13.647]<br />
are fairly close to each other</p>
<p>[00:33:15.780]<br />
and what the red line is is the actual original sample data.</p>
<p>[00:33:20.450]<br />
So these are the sample values</p>
<p>[00:33:22.250]<br />
and what we generally do want to see</p>
<p>[00:33:24.920]<br />
is that the estimators tend to bisect</p>
<p>[00:33:27.110]<br />
whether we have these highs and lows,</p>
<p>[00:33:29.280]<br />
the estimated values we should run through the middle.</p>
<p>[00:33:32.600]<br />
Also the estimator should follow the trend</p>
<p>[00:33:35.570]<br />
across the data.</p>
<p>[00:33:37.600]<br />
The nearest neighbor can be used as a validation tool.</p>
<p>[00:33:42.090]<br />
So if one of these estimators</p>
<p>[00:33:43.880]<br />
for instance was sitting quite high</p>
<p>[00:33:46.330]<br />
up here above their sniper line</p>
<p>[00:33:48.810]<br />
they would suggest that potentially where I over estimated</p>
<p>[00:33:50.850]<br />
and there might be some problems with our parameters</p>
<p>[00:33:54.040]<br />
so we would want to change that.</p>
<p>[00:33:56.810]<br />
Again we can export, we can copy this data</p>
<p>[00:34:00.537]<br />
and put it into a spreadsheet.</p>
<p>[00:34:02.160]<br />
Sometimes people have very specific formats</p>
<p>[00:34:06.650]<br />
they like to do their swath plots in.</p>
<p>[00:34:08.200]<br />
So that will bring all the data out into Excel.</p>
<p>[00:34:10.690]<br />
So you can do that yourself</p>
<p>[00:34:12.640]<br />
or we can just take the image</p>
<p>[00:34:14.830]<br />
and paste it directly into our report.</p>
<p>[00:34:21.780]<br />
Okay. If we want to report on our data,</p>
<p>[00:34:24.950]<br />
we create the report in the block model.</p>
<p>[00:34:28.400]<br />
And in this case, I’ve got just the basic report.</p>
<p>[00:34:33.710]<br />
That’s looking at the geology and the water table</p>
<p>[00:34:38.020]<br />
and it’s giving us the totals and the concentration.</p>
<p>[00:34:41.560]<br />
And this is by having to select these categorical columns</p>
<p>[00:34:45.400]<br />
and then we have our grade or value columns.</p>
<p>[00:34:48.030]<br />
So we need at least one categorical column</p>
<p>[00:34:50.820]<br />
and one to recreate a report</p>
<p>[00:34:53.310]<br />
but you can have as many different categories</p>
<p>[00:34:56.150]<br />
as you like within there.</p>
<p>[00:34:58.160]<br />
And these are also act like a pivot table almost</p>
<p>[00:35:01.030]<br />
so if I want it to show geology first</p>
<p>[00:35:04.370]<br />
and then want a table,</p>
<p>[00:35:05.760]<br />
I can split it up through there like that.</p>
<p>[00:35:08.220]<br />
We can apply a cutoff or we can have none as well.</p>
<p>[00:35:12.740]<br />
Another advantage within the block models (mumbles)</p>
<p>[00:35:20.090]<br />
is that we can create calculations and filters</p>
<p>[00:35:26.150]<br />
and we can get quite complex with these.</p>
<p>[00:35:30.040]<br />
We have a whole series of syntax helpers</p>
<p>[00:35:34.060]<br />
which are over here on the right.</p>
<p>[00:35:36.590]<br />
And then in middle here,</p>
<p>[00:35:37.750]<br />
it shows us all of our parameters</p>
<p>[00:35:39.620]<br />
that are available within the block model</p>
<p>[00:35:41.720]<br />
to use within a calculation.</p>
<p>[00:35:45.630]<br />
It’s also a nice little summary of the values</p>
<p>[00:35:48.800]<br />
within each estimator.</p>
<p>[00:35:52.030]<br />
Now you saw here for instance,</p>
<p>[00:35:53.900]<br />
I can see that I’ve got a couple of negatives</p>
<p>[00:35:56.797]<br />
in this ordinary krig without the variable orientation.</p>
<p>[00:36:00.270]<br />
So I’d want to have a look at those</p>
<p>[00:36:01.760]<br />
to see where they are and whether I need to remove them</p>
<p>[00:36:04.920]<br />
or do something about them.</p>
<p>[00:36:07.660]<br />
If I just unpin that for a second</p>
<p>[00:36:10.600]<br />
we can see, we can get quite complex</p>
<p>[00:36:13.020]<br />
we’ve got lots of if statements we can embed them.</p>
<p>[00:36:16.500]<br />
So what’s happening with this one</p>
<p>[00:36:18.060]<br />
is I’m trying to define porosity</p>
<p>[00:36:20.380]<br />
based on the rock type within the model.</p>
<p>[00:36:24.240]<br />
So I’ve defined, if we go back to here,</p>
<p>[00:36:28.740]<br />
I can look at the geology.</p>
<p>[00:36:30.820]<br />
So the geology is being coded within to the model.</p>
<p>[00:36:34.300]<br />
And if I stick on that block you can see here</p>
<p>[00:36:36.470]<br />
the geology model is till and the water table is saturated.</p>
<p>[00:36:41.060]<br />
So I can use any of those components (mumbles)</p>
<p>[00:36:48.723]<br />
so stop saying, «If the domain model is still,</p>
<p>[00:36:51.740]<br />
I’m going to assign a plus to the 8.25.»</p>
<p>[00:36:54.320]<br />
And that’s a fairly simple calculation</p>
<p>[00:36:56.300]<br />
but you can put in as many complex calculations as you wish.</p>
<p>[00:37:00.300]<br />
This one title concentration is saying that</p>
<p>[00:37:02.720]<br />
it must be within unconsolidated sediments.</p>
<p>[00:37:06.420]<br />
And what this one is doing is saying,</p>
<p>[00:37:07.960]<br />
what happens if I’ve got some blocks</p>
<p>[00:37:09.580]<br />
that haven’t been given a value.</p>
<p>[00:37:12.140]<br />
So I can say that if there is no value,</p>
<p>[00:37:15.059]<br />
I’m going to assign a very low one</p>
<p>[00:37:17.570]<br />
but otherwise use the value if it’s greater than zero,</p>
<p>[00:37:21.270]<br />
use the value that’s there.</p>
<p>[00:37:22.510]<br />
So that’s one way of getting rid of negatives</p>
<p>[00:37:24.440]<br />
if you know that there’s only a</p>
<p>[00:37:26.330]<br />
couple of isolated ones there.</p>
<p>[00:37:30.680]<br />
And then this one is an average distance.</p>
<p>[00:37:32.530]<br />
So I store the average distance within the model.</p>
<p>[00:37:35.980]<br />
So I’m using this as a guide for classification.</p>
<p>[00:37:39.260]<br />
So stop saying if the average distance</p>
<p>[00:37:40.940]<br />
to the samples is less than 500 meters,</p>
<p>[00:37:43.500]<br />
I’ll make it measured less than 1,000 indicated</p>
<p>[00:37:46.621]<br />
less than 1500.</p>
<p>[00:37:48.970]<br />
The beauty of any calculation, numeric or categorical</p>
<p>[00:37:52.630]<br />
is that it’s available instantly to visualize.</p>
<p>[00:37:55.810]<br />
So we can visualize this confidence category.</p>
<p>[00:37:59.410]<br />
So you can see that there,</p>
<p>[00:38:00.610]<br />
I’ve just got indicated in third</p>
<p>[00:38:03.380]<br />
and a little bit of measured there as well.</p>
<p>[00:38:06.800]<br />
Let’s split that out. (mumbles)</p>
<p>[00:38:10.980]<br />
So we can see here,</p>
<p>[00:38:12.360]<br />
that’s based on the data and the data spacing.</p>
<p>[00:38:17.050]<br />
We might not use those values directly to classify</p>
<p>[00:38:19.860]<br />
but we can use them as a visual guide</p>
<p>[00:38:22.240]<br />
if we wanted to draw shapes to classify certain areas.</p>
<p>[00:38:28.580]<br />
So once we’ve done that kind of thing</p>
<p>[00:38:31.100]<br />
we can build another report based of that calculation.</p>
<p>[00:38:36.460]<br />
So in this case we’re using that calculated field</p>
<p>[00:38:40.060]<br />
as part of that categorical information.</p>
<p>[00:38:43.920]<br />
And so now we’ve got classification,</p>
<p>[00:38:46.180]<br />
the geology and the water table in the report.</p>
<p>[00:38:49.630]<br />
And again that’s saved automatically and it can be exported.</p>
<p>[00:38:53.890]<br />
We can copy it into an Excel Spreadsheet</p>
<p>[00:38:57.790]<br />
or you can just export it as an Excel Spreadsheet directly.</p>
<p>[00:39:05.930]<br />
Okay. So that’s the main usage</p>
<p>[00:39:10.240]<br />
of the Contaminants Extension.</p>
<p>[00:39:12.160]<br />
And I’ll just pass right back again to Aaron to finish it.</p>
<p>[00:39:16.290]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Fantastic. Thanks Steve.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:39:20.160]<br />
We have time now for questions as well.</p>
<p>[00:39:23.670]<br />
So if you have them already,</p>
<p>[00:39:25.360]<br />
please feel free to put a question in the question box</p>
<p>[00:39:28.330]<br />
or you can raise your hand as well</p>
<p>[00:39:30.320]<br />
and we can unmute you</p>
<p>[00:39:31.670]<br />
and you can verbally ask the question as well.</p>
<p>[00:39:35.650]<br />
While we give people a chance to do that,</p>
<p>[00:39:39.289]<br />
beyond today’s meeting there’s always support for Leapfrog</p>
<p>[00:39:43.360]<br />
and the Contamination Extension</p>
<p>[00:39:44.880]<br />
by contacting us at [email protected]</p>
<p>[00:39:48.910]<br />
or by giving our support line a phone call.</p>
<p>[00:39:52.480]<br />
After today’s session as well,</p>
<p>[00:39:54.852]<br />
we’d love to give the extension of Leapfrog a go.</p>
<p>[00:39:59.620]<br />
We have a trial available</p>
<p>[00:40:00.790]<br />
that you can sign up for on our website</p>
<p>[00:40:02.480]<br />
on the Leapfrog Works products page</p>
<p>[00:40:04.450]<br />
which you can see the URL therefore.</p>
<p>[00:40:06.690]<br />
We’ll also be sending an email out after this webinar</p>
<p>[00:40:10.710]<br />
with the recording for today</p>
<p>[00:40:12.410]<br />
and also a link for a four-part technical series</p>
<p>[00:40:16.180]<br />
on the Contamination Extension</p>
<p>[00:40:17.840]<br />
which will take you through step-by-step</p>
<p>[00:40:20.330]<br />
how to build the model that Steve has shown you</p>
<p>[00:40:23.190]<br />
from scratch and the process behind that.</p>
<p>[00:40:27.440]<br />
And includes the data there as well.</p>
<p>[00:40:28.830]<br />
So that’s a fantastic resource we’ll be sending to you.</p>
<p>[00:40:31.860]<br />
We also have online learning content</p>
<p>[00:40:34.490]<br />
at my.seequent.com/learning.</p>
<p>[00:40:37.500]<br />
And we also have the</p>
<p>[00:40:38.830]<br />
Leapfrog Works and Contaminants Extension help pages</p>
<p>[00:40:41.910]<br />
will look out at them later.</p>
<p>[00:40:43.400]<br />
And that’s a really useful resource</p>
<p>[00:40:44.850]<br />
if you have any questions.</p>
<p>[00:40:51.770]<br />
So yeah like I said, if you have questions,</p>
<p>[00:40:54.510]<br />
please put them up or raise your hand.</p>
<p>[00:40:57.170]<br />
One of the questions that we have so far is</p>
<p>[00:41:01.392]<br />
so often contaminants sources</p>
<p>[00:41:03.840]<br />
are at the surface propagating down</p>
<p>[00:41:05.910]<br />
into the soil or groundwater</p>
<p>[00:41:09.872]<br />
and how would you go about modeling a change</p>
<p>[00:41:12.800]<br />
in the directional trend</p>
<p>[00:41:14.240]<br />
I guess of the plume going from vertical</p>
<p>[00:41:16.620]<br />
as it propagates down to spreading out horizontally.</p>
<p>[00:41:20.075]<br />
I guess once it’s an aquifer or an aquatar.</p>
<p>[00:41:24.710]<br />
So can you answer that for us please Steve?</p>
<p>[00:41:27.230]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Yep so everything about the Contaminants Extension<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:41:30.750]<br />
is all about how we set up our domains</p>
<p>[00:41:33.360]<br />
at the very beginning.</p>
<p>[00:41:34.770]<br />
So it does need a physical three-dimensional representation</p>
<p>[00:41:39.056]<br />
of each domain.</p>
<p>[00:41:40.690]<br />
So the way we would do that would</p>
<p>[00:41:42.310]<br />
we would need acquifer surface and the soil.</p>
<p>[00:41:46.414]<br />
We would have one domain and say that</p>
<p>[00:41:49.390]<br />
then you’d model your aquifer as a horizontal layer.</p>
<p>[00:41:54.410]<br />
So you would set up two estimation folders,</p>
<p>[00:41:57.120]<br />
one for the soil, one for the aquifer.</p>
<p>[00:41:59.410]<br />
And then within the soil one,</p>
<p>[00:42:00.630]<br />
you could make your orientation vertical</p>
<p>[00:42:03.420]<br />
and within the aquifer you can change it to horizontal.</p>
<p>[00:42:06.850]<br />
And then you get…</p>
<p>[00:42:07.770]<br />
You’re able to join these things together</p>
<p>[00:42:10.120]<br />
using the calculation so that you can visualize</p>
<p>[00:42:13.240]<br />
both domains together in the block model.</p>
<p>[00:42:16.200]<br />
So you only need one block model</p>
<p>[00:42:17.920]<br />
but you can have multiple domains</p>
<p>[00:42:19.890]<br />
which will get twined together.</p>
<p>[00:42:21.950]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Fantastic. Thank you Steve.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:42:24.770]<br />
Another question here is,</p>
<p>[00:42:26.600]<br />
were any of those calculations</p>
<p>[00:42:28.180]<br />
automatically populated by Leapfrog</p>
<p>[00:42:30.270]<br />
or did you make each of them?</p>
<p>[00:42:32.040]<br />
If you made them, just to be clear,</p>
<p>[00:42:33.530]<br />
did you make them to remove the negative values</p>
<p>[00:42:35.960]<br />
generated by the kriging?</p>
<p>[00:42:38.045]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />No, they’re not generated automatically.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:42:40.530]<br />
So I created all of those calculations</p>
<p>[00:42:43.450]<br />
and you don’t have to force removal of the negatives.</p>
<p>[00:42:48.510]<br />
It really comes down to first</p>
<p>[00:42:50.860]<br />
realizing that negative values exist.</p>
<p>[00:42:53.420]<br />
And then we would look at them in the 3D scene</p>
<p>[00:42:55.450]<br />
to see where they are.</p>
<p>[00:42:56.820]<br />
Now, if they’re only the isolated block around the edges</p>
<p>[00:43:00.260]<br />
quite a bit away from your data,</p>
<p>[00:43:03.010]<br />
then they might not be significant.</p>
<p>[00:43:05.060]<br />
So then you could use the calculation to zero them out.</p>
<p>[00:43:09.690]<br />
Sometimes you get a situation</p>
<p>[00:43:11.260]<br />
where you can get negative grades</p>
<p>[00:43:13.410]<br />
right within the main part of your data.</p>
<p>[00:43:16.387]<br />
And this is the thing called a kriging et cetera.</p>
<p>[00:43:21.763]<br />
and the kriging process can cause this.</p>
<p>[00:43:24.550]<br />
So in that case you would have to</p>
<p>[00:43:26.100]<br />
change your search selection.</p>
<p>[00:43:28.510]<br />
So you might have to decrease the maximum number of samples.</p>
<p>[00:43:31.750]<br />
You might need to tweak the variogram a little bit</p>
<p>[00:43:34.340]<br />
and then you should be able to get</p>
<p>[00:43:35.830]<br />
reduce the impact of negative weights</p>
<p>[00:43:38.219]<br />
So that you won’t get negative grades.</p>
<p>[00:43:41.450]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />To add to that first part of the question as well.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:43:43.942]<br />
Once you have created your calculations,</p>
<p>[00:43:47.230]<br />
you can actually go in and export them out</p>
<p>[00:43:51.760]<br />
to use in other projects as well.</p>
<p>[00:43:53.410]<br />
So kind of… Where are we?</p>
<p>[00:43:58.490]<br />
Calculations are cool.</p>
<p>[00:43:59.860]<br />
So calculations and filters.</p>
<p>[00:44:01.490]<br />
So in this calculations tab here,</p>
<p>[00:44:03.780]<br />
you can see we have this option here</p>
<p>[00:44:05.490]<br />
for importing or indeed for exporting here as well.</p>
<p>[00:44:08.840]<br />
So once you set it at once,</p>
<p>[00:44:10.440]<br />
you can reuse that in other projects.</p>
<p>[00:44:12.620]<br />
So it’s very easy to transfer these across projects</p>
<p>[00:44:16.260]<br />
maybe change some of the variable names.</p>
<p>[00:44:18.260]<br />
If you need to, it’s different under different projects.</p>
<p>[00:44:20.967]<br />
And it’s once you’ve set at once</p>
<p>[00:44:22.410]<br />
it’s very easy to use those again</p>
<p>[00:44:24.167]<br />
and that’s a files, it’s a calculation file</p>
<p>[00:44:26.380]<br />
that you can send to other people in your company as well.</p>
<p>[00:44:28.390]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />So if you don’t have the same variable names,<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:44:31.830]<br />
what it does,</p>
<p>[00:44:32.663]<br />
it just highlights them with a little red,</p>
<p>[00:44:34.430]<br />
underline and then you just replace it</p>
<p>[00:44:36.880]<br />
with the correct variable name.</p>
<p>[00:44:38.340]<br />
So yeah easy to set things up</p>
<p>[00:44:41.890]<br />
especially if you’re using the same format</p>
<p>[00:44:43.545]<br />
of block model over a whole project area</p>
<p>[00:44:46.400]<br />
for different block models.</p>
<p>[00:44:49.020]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Fantastic. And yeah as a reminder,<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:44:51.230]<br />
if you do have questions,</p>
<p>[00:44:52.150]<br />
please put them in the chat or raise your hand.</p>
<p>[00:44:55.140]<br />
We have another question here.</p>
<p>[00:44:56.220]<br />
Can you run the steps to include the 95 percentile?</p>
<p>[00:45:01.562]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Not directly,<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:45:04.331]<br />
actually we can get…</p>
<p>[00:45:07.090]<br />
We do do log probability plots.</p>
<p>[00:45:09.600]<br />
So (mumbles).</p>
<p>[00:45:13.350]<br />
So if I go back up to this one here yet</p>
<p>[00:45:24.680]<br />
so my daily statistics of strike first up</p>
<p>[00:45:27.940]<br />
get a histogram,</p>
<p>[00:45:30.169]<br />
but I can change this to a low probability plot.</p>
<p>[00:45:32.710]<br />
So then I’ll be able to see where the 95th percentile is.</p>
<p>[00:45:36.890]<br />
Is down there.</p>
<p>[00:45:38.570]<br />
So the 95th percentile is up through there like that.</p>
<p>[00:45:42.622]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Awesome. Thanks Steve.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:45:45.850]<br />
So can you please show</p>
<p>[00:45:48.310]<br />
the contamination plume advance over time?</p>
<p>[00:45:50.790]<br />
And can you show the database with the time element column?</p>
<p>[00:45:54.990]<br />
So we don’t have any time or…</p>
<p>[00:45:58.534]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Sorry.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:46:00.184]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />monitoring data set up for this project<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:46:03.321]<br />
but you could certainly do that.</p>
<p>[00:46:05.140]<br />
So you could just set up your first one,</p>
<p>[00:46:07.555]<br />
make a copy of it and then change the data to the new,</p>
<p>[00:46:10.730]<br />
but the next monitoring event.</p>
<p>[00:46:11.920]<br />
So if each monitoring event,</p>
<p>[00:46:13.654]<br />
you’re just using that separate data</p>
<p>[00:46:16.400]<br />
in the same estimations, in the same block models</p>
<p>[00:46:20.900]<br />
and also in with the contaminants kit.</p>
<p>[00:46:23.250]<br />
If you go into the points folder,</p>
<p>[00:46:25.335]<br />
you do have this option for</p>
<p>[00:46:26.900]<br />
import time dependent points as well.</p>
<p>[00:46:28.830]<br />
So you can import a point cloud</p>
<p>[00:46:30.070]<br />
that has the time data or attribute to it.</p>
<p>[00:46:33.370]<br />
You can actually filter that.</p>
<p>[00:46:34.390]<br />
So visually you can use a filter</p>
<p>[00:46:36.040]<br />
to see the data at a certain point in time</p>
<p>[00:46:38.340]<br />
but you could then actually</p>
<p>[00:46:40.050]<br />
if you wanted to, you could set up a workflow</p>
<p>[00:46:41.960]<br />
where you set filters up</p>
<p>[00:46:43.550]<br />
to use that point data in your block model</p>
<p>[00:46:46.630]<br />
and just filter it based on time as well.</p>
<p>[00:46:49.650]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />So when you create a estimator,<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:46:53.550]<br />
so this one here,</p>
<p>[00:46:59.390]<br />
you can see here you can apply a filter to your data.</p>
<p>[00:47:02.420]<br />
So yes, if you divided up</p>
<p>[00:47:05.030]<br />
created pre filters based on the time,</p>
<p>[00:47:07.660]<br />
you can run a series of block models.</p>
<p>[00:47:13.090]<br />
Well actually you could probably do it</p>
<p>[00:47:14.700]<br />
within the single block model.</p>
<p>[00:47:16.090]<br />
So you would just set up</p>
<p>[00:47:17.440]<br />
a different estimator for each time period,</p>
<p>[00:47:20.020]<br />
estimate them all and put them all,</p>
<p>[00:47:21.810]<br />
evaluate them all into the same block model</p>
<p>[00:47:24.260]<br />
and then you could kind of look at them and how it changes.</p>
<p>[00:47:28.070]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Yeah. Awesome.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:47:29.640]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />That’d be a few different ways<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:47:30.790]<br />
of trying to do that.</p>
<p>[00:47:35.420]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />We have a question here<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:47:36.770]<br />
which is, when should you use the different</p>
<p>[00:47:41.750]<br />
or when should you use the different estimation methods</p>
<p>[00:47:45.190]<br />
or I guess?</p>
<p>[00:47:46.478]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Guess so. It really comes down<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:47:48.860]<br />
to personal choice a little bit</p>
<p>[00:47:51.810]<br />
but generally, we do need a certain degree</p>
<p>[00:47:56.270]<br />
of the amount of data to do kriging</p>
<p>[00:47:58.590]<br />
so that we can get a variable variogram</p>
<p>[00:48:01.420]<br />
So early stages,</p>
<p>[00:48:02.670]<br />
we might only be able to do inverse distance</p>
<p>[00:48:06.510]<br />
whether it’s inverse distance squared or cubed et cetera.</p>
<p>[00:48:10.300]<br />
It’s again personal preference.</p>
<p>[00:48:13.517]<br />
Generally though, if we wanted to get</p>
<p>[00:48:15.280]<br />
quite a localized estimate,</p>
<p>[00:48:17.780]<br />
we may use inverse distance cubed,</p>
<p>[00:48:19.860]<br />
a more general one the inverse distance squared.</p>
<p>[00:48:23.580]<br />
And now if we can start to get a variogram out</p>
<p>[00:48:25.960]<br />
it’s always best to use kriging if possible</p>
<p>[00:48:28.777]<br />
it does tend.</p>
<p>[00:48:29.970]<br />
You find that, especially with sparse data,</p>
<p>[00:48:32.760]<br />
you won’t get too much,</p>
<p>[00:48:34.270]<br />
the results shouldn’t be too different from each other</p>
<p>[00:48:36.930]<br />
but kriging does tend to work with it better.</p>
<p>[00:48:41.150]<br />
The RBF, I don’t use it much myself</p>
<p>[00:48:44.173]<br />
but it can be quite useful to get early stage global idea</p>
<p>[00:48:49.590]<br />
of how things are going.</p>
<p>[00:48:50.790]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Yeah. So when you have sparse<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:48:51.847]<br />
so not much data can be useful in those initial stages</p>
<p>[00:48:55.120]<br />
maybe as pilot or desk study phase or something.</p>
<p>[00:48:57.930]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Yeah. Yeah.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:48:58.871]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />For some case (mumbles).<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:49:02.110]<br />
Another question here,</p>
<p>[00:49:03.020]<br />
can you please explain the hard and soft boundaries</p>
<p>[00:49:05.450]<br />
that appeared in the box just before</p>
<p>[00:49:08.280]<br />
so in that initial estimation and this one here.</p>
<p>[00:49:13.550]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />So what this is,<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:49:15.190]<br />
is that in this case it’s not</p>
<p>[00:49:17.408]<br />
or that sort of relevant.</p>
<p>[00:49:18.990]<br />
There are only two samples outside our domain.</p>
<p>[00:49:21.930]<br />
So if I bring back up my domain boundary,</p>
<p>[00:49:25.900]<br />
let’s do that, okay.</p>
<p>[00:49:31.090]<br />
So hard and soft basically</p>
<p>[00:49:32.982]<br />
is what happens around the boundary of the domain itself.</p>
<p>[00:49:37.950]<br />
How do I treat the samples?</p>
<p>[00:49:40.110]<br />
So a hard boundary in 99% of the time,</p>
<p>[00:49:43.720]<br />
you will use a hard boundary.</p>
<p>[00:49:45.430]<br />
Is that you saying,</p>
<p>[00:49:47.180]<br />
basically I’m only looking at samples</p>
<p>[00:49:48.930]<br />
within the domain itself,</p>
<p>[00:49:50.300]<br />
and I don’t care about anything outside it.</p>
<p>[00:49:53.630]<br />
The instance where you may use a soft boundary</p>
<p>[00:49:56.030]<br />
is if there’s a good directional change,</p>
<p>[00:49:59.660]<br />
say the concentration can’t be limited directly</p>
<p>[00:50:02.630]<br />
to one of the main or another.</p>
<p>[00:50:03.463]<br />
And there’s a good directional change across that boundary.</p>
<p>[00:50:07.200]<br />
By using a soft boundary, we can define,</p>
<p>[00:50:11.222]<br />
we could say 20 meters.</p>
<p>[00:50:13.600]<br />
And what that would do is we’ll look at for samples,</p>
<p>[00:50:16.660]<br />
20 meters beyond the boundary in all directions.</p>
<p>[00:50:19.650]<br />
And we’ll use those as part of the sample selection.</p>
<p>[00:50:23.350]<br />
You still only estimate within the domain,</p>
<p>[00:50:27.420]<br />
but around the edges you use samples</p>
<p>[00:50:29.860]<br />
that are just slightly outside.</p>
<p>[00:50:31.730]<br />
So that’s the main difference between hard and soft.</p>
<p>[00:50:35.090]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />And I guess going back<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:50:36.180]<br />
to a question and an answer</p>
<p>[00:50:37.330]<br />
that was before around using multiple domains</p>
<p>[00:50:40.260]<br />
to kind of handle the vertical and then horizontal</p>
<p>[00:50:42.900]<br />
would it be fair to say</p>
<p>[00:50:43.733]<br />
that maybe you’d use the soft boundary</p>
<p>[00:50:45.700]<br />
in that first one to accommodate the values near</p>
<p>[00:50:50.130]<br />
really that domain boundary into the horizontal?</p>
<p>[00:50:52.290]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />That’s right.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:50:53.250]<br />
And I mean, one of the beauties of a Leapfrog Works,</p>
<p>[00:50:57.960]<br />
the Contaminants et cetera,</p>
<p>[00:50:59.210]<br />
is that it’s so easy to test things.</p>
<p>[00:51:02.020]<br />
So once you’ve got one of these folders set up</p>
<p>[00:51:06.060]<br />
if you want to try something a little bit different,</p>
<p>[00:51:08.420]<br />
you just copy the entire folder</p>
<p>[00:51:11.470]<br />
and you might pick a slightly different domain</p>
<p>[00:51:14.630]<br />
or if you’re going to do it on a different element,</p>
<p>[00:51:16.910]<br />
choose it there.</p>
<p>[00:51:18.310]<br />
And let’s just copy. That’s fine.</p>
<p>[00:51:20.360]<br />
And it pre copies basically any very variables</p>
<p>[00:51:22.923]<br />
that you might have done or you estimated is a set up.</p>
<p>[00:51:25.720]<br />
And then you can just go back into that one and say,</p>
<p>[00:51:28.940]<br />
I want to try something that’s little bit different</p>
<p>[00:51:31.237]<br />
with this kriging set up.</p>
<p>[00:51:33.150]<br />
So you might try a different distance,</p>
<p>[00:51:35.710]<br />
some different samples et cetera.</p>
<p>[00:51:38.050]<br />
So then you would just run them,</p>
<p>[00:51:39.645]<br />
evaluate that into your bottle</p>
<p>[00:51:41.740]<br />
and look at it on a swath plot</p>
<p>[00:51:43.760]<br />
to see what the difference was.</p>
<p>[00:51:45.820]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Awesome. Thanks Steve.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:51:48.370]<br />
I think you touched on maybe this question a little bit,</p>
<p>[00:51:51.040]<br />
but is there an output or evaluation or really…</p>
<p>[00:51:56.150]<br />
How can you show uncertainty of the block model?</p>
<p>[00:51:58.760]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Okay. So there are lots of different ways<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:52:01.650]<br />
of trying to look at uncertainty.</p>
<p>[00:52:03.743]<br />
The simplest way is just looking at data spacing.</p>
<p>[00:52:07.469]<br />
And that’s a little bit what we’ve tried to do</p>
<p>[00:52:10.240]<br />
with the calculation field before.</p>
<p>[00:52:13.860]<br />
So I open up my calculations again.</p>
<p>[00:52:16.550]<br />
So we stored the average distance to the samples</p>
<p>[00:52:20.430]<br />
and that was done.</p>
<p>[00:52:23.690]<br />
I’m here in the outputs</p>
<p>[00:52:25.460]<br />
so I stored the average distance sample.</p>
<p>[00:52:27.530]<br />
You can also store the minimum distance</p>
<p>[00:52:29.720]<br />
to the closest sample.</p>
<p>[00:52:31.600]<br />
And then again by making up</p>
<p>[00:52:33.960]<br />
some kind of a calculation like this,</p>
<p>[00:52:38.130]<br />
if that’s not working, then I can change.</p>
<p>[00:52:40.370]<br />
And I could change that to 800, 1200 et cetera</p>
<p>[00:52:44.240]<br />
until it sort of makes sense.</p>
<p>[00:52:46.210]<br />
So we can use (mumbles).</p>
<p>[00:52:50.040]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Yeah.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:52:50.873]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />We can use distance buffers,<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:52:52.060]<br />
like let’s add on</p>
<p>[00:52:53.900]<br />
Contaminants Extension functionality within works</p>
<p>[00:52:56.530]<br />
and that might help you as well</p>
<p>[00:52:58.440]<br />
decide on what kind of values to use</p>
<p>[00:53:00.630]<br />
within these calculations.</p>
<p>[00:53:02.300]<br />
Classification, if you’re using kriging,</p>
<p>[00:53:06.046]<br />
if you have used directly kriging variance</p>
<p>[00:53:09.270]<br />
as it often quite nicely to use spacing as well.</p>
<p>[00:53:12.980]<br />
So there’s lots of different ways,</p>
<p>[00:53:15.220]<br />
but in more of the LT user.</p>
<p>[00:53:17.520]<br />
<encoded_tag_open />v Aaron<encoded_tag_closed />Fantastic. Well thank you so much Steve.<encoded_tag_open />/v<encoded_tag_closed /></p>
<p>[00:53:20.317]<br />
We’re coming to the end of…</p>
<p>[00:53:22.107]<br />
We had a lot of time now.</p>
<p>[00:53:23.830]<br />
Thank you, everyone who joined</p>
<p>[00:53:25.470]<br />
and particularly if you asked a question.</p>
<p>[00:53:27.902]<br />
If we didn’t get to your question</p>
<p>[00:53:29.800]<br />
or if you think of one after today’s session,</p>
<p>[00:53:32.360]<br />
then please feel free to reach out</p>
<p>[00:53:35.990]<br />
at the [email protected].</p>
<p>[00:53:38.100]<br />
And if there are anything we couldn’t get though today,</p>
<p>[00:53:40.480]<br />
we’ll also reach out directly to people.</p>
<p>[00:53:44.720]<br />
As I said, you will be receiving a link after today</p>
<p>[00:53:49.450]<br />
with the recording as well as</p>
<p>[00:53:51.183]<br />
the link for that technical series,</p>
<p>[00:53:53.700]<br />
which is a really good resource to go through yourself</p>
<p>[00:53:56.960]<br />
using the Contaminant Extension.</p>
<p>[00:53:59.160]<br />
And if you don’t already have access to it,</p>
<p>[00:54:00.790]<br />
we can give you a trial of it as well.</p>
<p>[00:54:03.410]<br />
So you can give it a go.</p>
<p>[00:54:07.840]<br />
Thank you so much for joining today’s webinar.</p>
<p>[00:54:09.960]<br />
We really hope you enjoyed it</p>
<p>[00:54:11.450]<br />
and have a fantastic rest of your day.</p>
<wpml_invalid_tag original=»PHA+» />[00:54:13.590]<br />
<encoded_tag_open />v Steve<encoded_tag_closed />Thank you very much.<encoded_tag_open />/v<encoded_tag_closed />